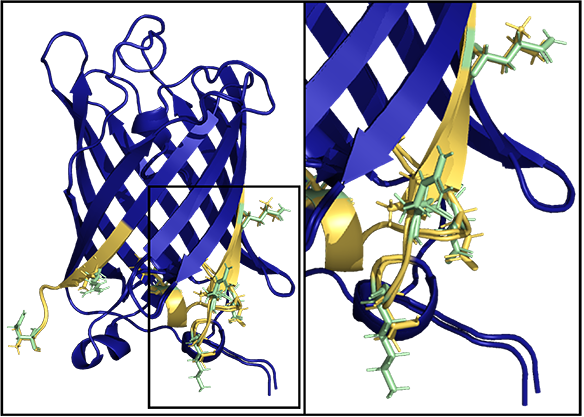
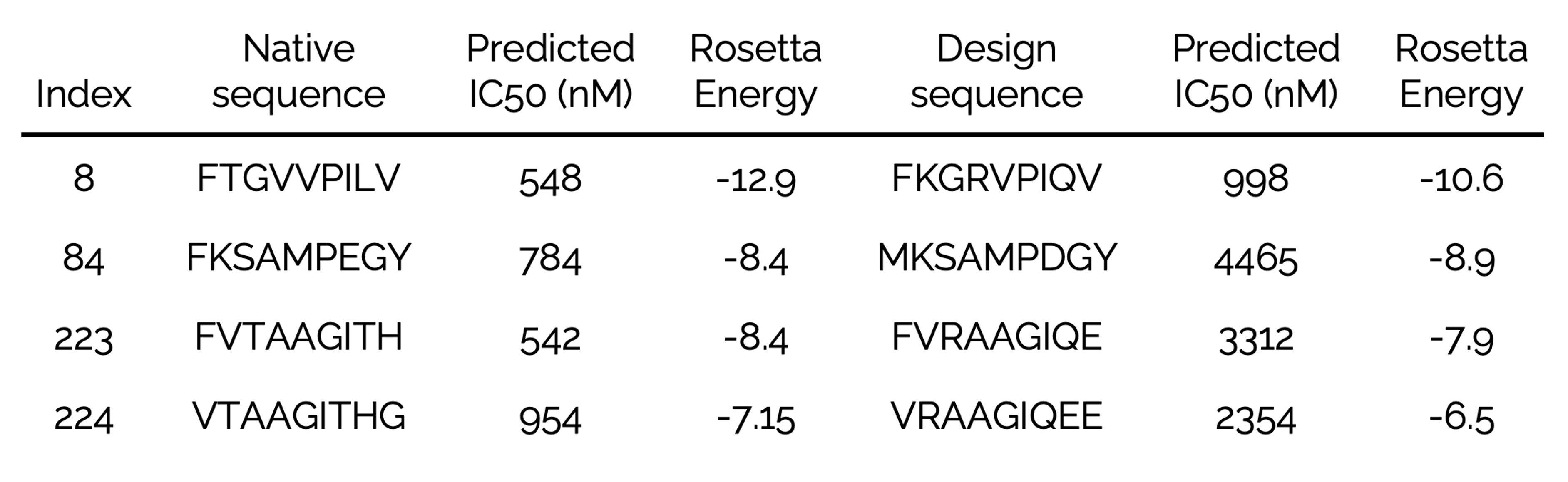
As proof-of-concept, Green Fluorescent Protein (GFP) was redesigned to deimmunize. GFP, shown in blue, with immunodominant epitopes of shown in yellow. Mutations in green were selected by ML-guided, Rosetta-based epitope redesign. As shown in Removing T-cell epitopes with computational protein design. King C, et al. Proc Natl Acad Sci U S A. 2014
Reducing Immune Responses to Protein Drugs
Immunogenicity is a significant challenge for biologics
Patients are often treated with a therapeutic protein to which they’ve had no prior exposure, perhaps because they lack the protein naturally, or it’s a significantly engineered version. They may have inherited a damaged or missing gene and do not produce the protein naturally and a replacement protein is therapeutic. Or they may suffer from an illness that can be effectively treated with a non-human protein.
Perceived as foreign, the body mounts an immune response against the therapeutic protein. Anti-drug antibodies (ADAs) form and bind to the therapeutic, clearing from the bloodstream and significantly reducing its efficacy. In rarer cases, the patient’s immune response may be strong enough to cause a serious allergic reaction, possibly including anaphylaxis or even cardiac arrest. About a quarter of all approved protein biologics are known to have measurable immunogenicity (>5% of patients with signs of an immune response).
T-cell Epitopes
T-cell epitopes – the regions of a protein that can trigger these immune responses via the MHC complex – can be identified both through empirical in vitro assays and via computational prediction. The rational modification of these epitopes has been difficult, because of the complexity of protein structure. Using our unique platform, Cyrus can use software to redesign and rebuild proteins, removing immune epitopes while maintaining key structural features for the protein’s normal function.
To replace an immune epitope of 15 amino acids requires the evaluation of up to 1019 different amino acid combinations, an enormous number and something clearly impossible to tackle experimentally. Most of these substitutions will destabilize the protein and prove useless. That’s where a computational approach combining Rosetta with ML comes into play. Because our algorithms permit the simultaneous evaluation of more than 109 substitutions in hours, identifying productive replacements becomes a tractable problem.
Cyrus combines Rosetta and ML-based epitope predictions for deimmunization. Rosetta is excellent for selecting mutations that retain structure/function and epitope predictions will bias mutation-selection towards favorable mutations that are not predicted to have an immune response. This allows Cyrus to develop proteins that are less likely to trigger ADAs while retaining therapeutic function.
INTERESTED IN
PARTNERING WITH US ON
PROTEIN THERAPEUTICS?